hd_plot_model_summary() plots the number of features and the number of top
features (feature importance > user defined threshold) for each disease in a barplot.
It also plots the upset plot of the top or all features, as well as a summary line plot
of the model performance metrics.
Usage
hd_plot_model_summary(
model_results,
importance = 0.5,
class_palette = NULL,
upset_top_features = FALSE
)Arguments
- model_results
A list of binary classification model results. It should be a list of objects created by
hd_model_rreg(),hd_model_rf()orhd_model_lr()with the classes as names. See the examples for more details.- importance
The importance threshold to consider a feature as top. Default is 0.5.
- class_palette
The color palette for the classes. If it is a character, it should be one of the palettes from
hd_palettes(). Default is NULL.- upset_top_features
Whether to plot the upset plot for the top features or all features. Default is FALSE (all features).
Examples
# Initialize an HDAnalyzeR object with only a subset of the predictors
hd_object <- hd_initialize(example_data, example_metadata)
# Split the data into training and test sets
hd_split <- hd_split_data(hd_object, variable = "Disease")
#> Warning: Too little data to stratify.
#> • Resampling will be unstratified.
# Run the regularized regression model pipeline
model_results_aml <- hd_model_rreg(hd_split,
variable = "Disease",
case = "AML",
grid_size = 2,
cv_sets = 2,
verbose = FALSE)
#> The groups in the train set are balanced. If you do not want to balance the groups, set `balance_groups = FALSE`.
model_results_cll <- hd_model_rreg(hd_split,
variable = "Disease",
case = "CLL",
grid_size = 2,
cv_sets = 2,
verbose = FALSE)
#> The groups in the train set are balanced. If you do not want to balance the groups, set `balance_groups = FALSE`.
model_results_myel <- hd_model_rreg(hd_split,
variable = "Disease",
case = "MYEL",
grid_size = 2,
cv_sets = 2,
verbose = FALSE)
#> The groups in the train set are balanced. If you do not want to balance the groups, set `balance_groups = FALSE`.
model_results_lungc <- hd_model_rreg(hd_split,
variable = "Disease",
case = "LUNGC",
grid_size = 2,
cv_sets = 2,
verbose = FALSE)
#> The groups in the train set are balanced. If you do not want to balance the groups, set `balance_groups = FALSE`.
model_results_gliom <- hd_model_rreg(hd_split,
variable = "Disease",
case = "GLIOM",
grid_size = 2,
cv_sets = 2,
verbose = FALSE)
#> The groups in the train set are balanced. If you do not want to balance the groups, set `balance_groups = FALSE`.
res <- list("AML" = model_results_aml,
"LUNGC" = model_results_lungc,
"CLL" = model_results_cll,
"MYEL" = model_results_myel,
"GLIOM" = model_results_gliom)
# Plot summary visualizations
hd_plot_model_summary(res, class_palette = "cancers12")
#> $features_barplot
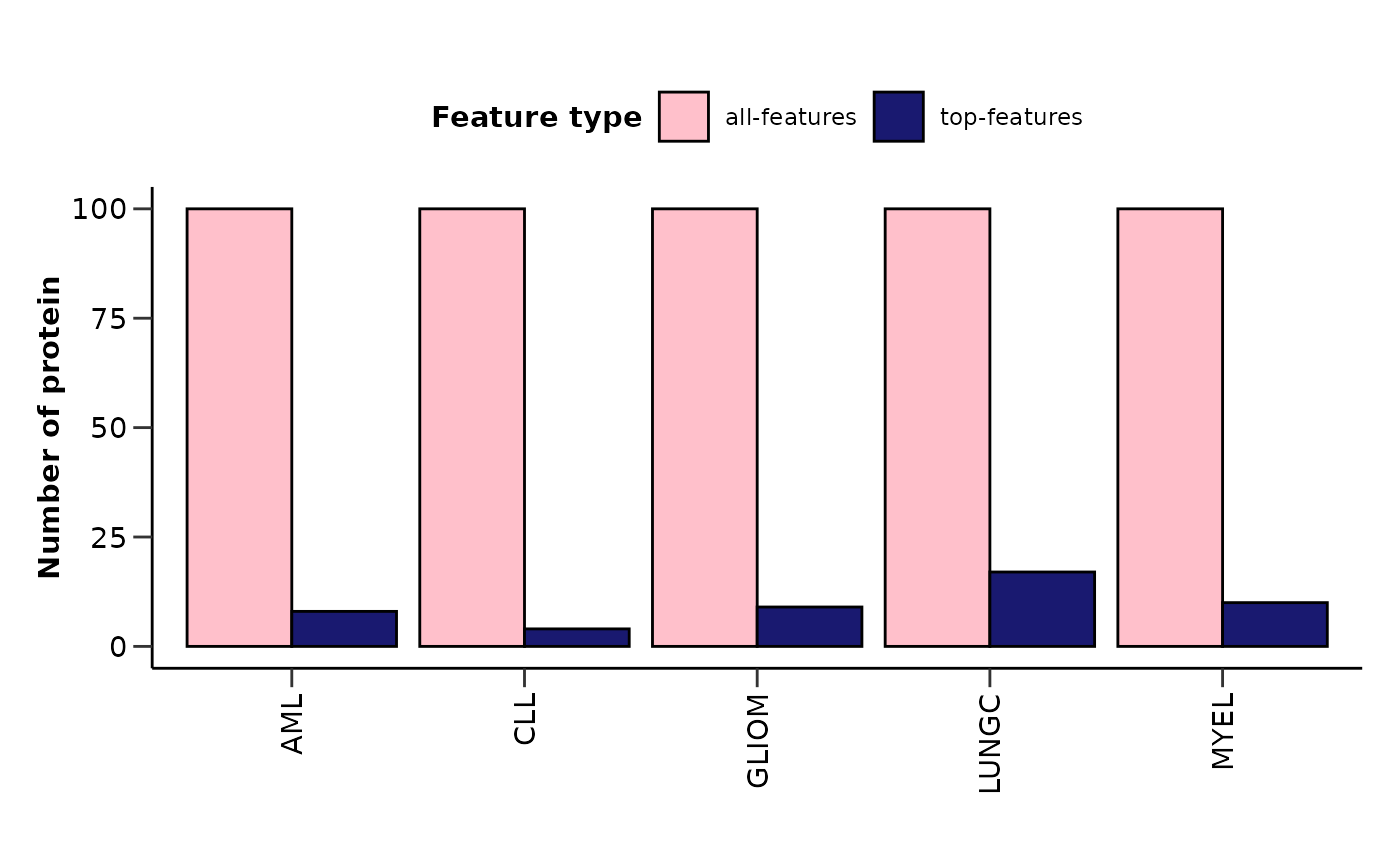 #>
#> $metrics_barplot
#> Ignoring unknown labels:
#> • colour : "Metric"
#>
#> $metrics_barplot
#> Ignoring unknown labels:
#> • colour : "Metric"
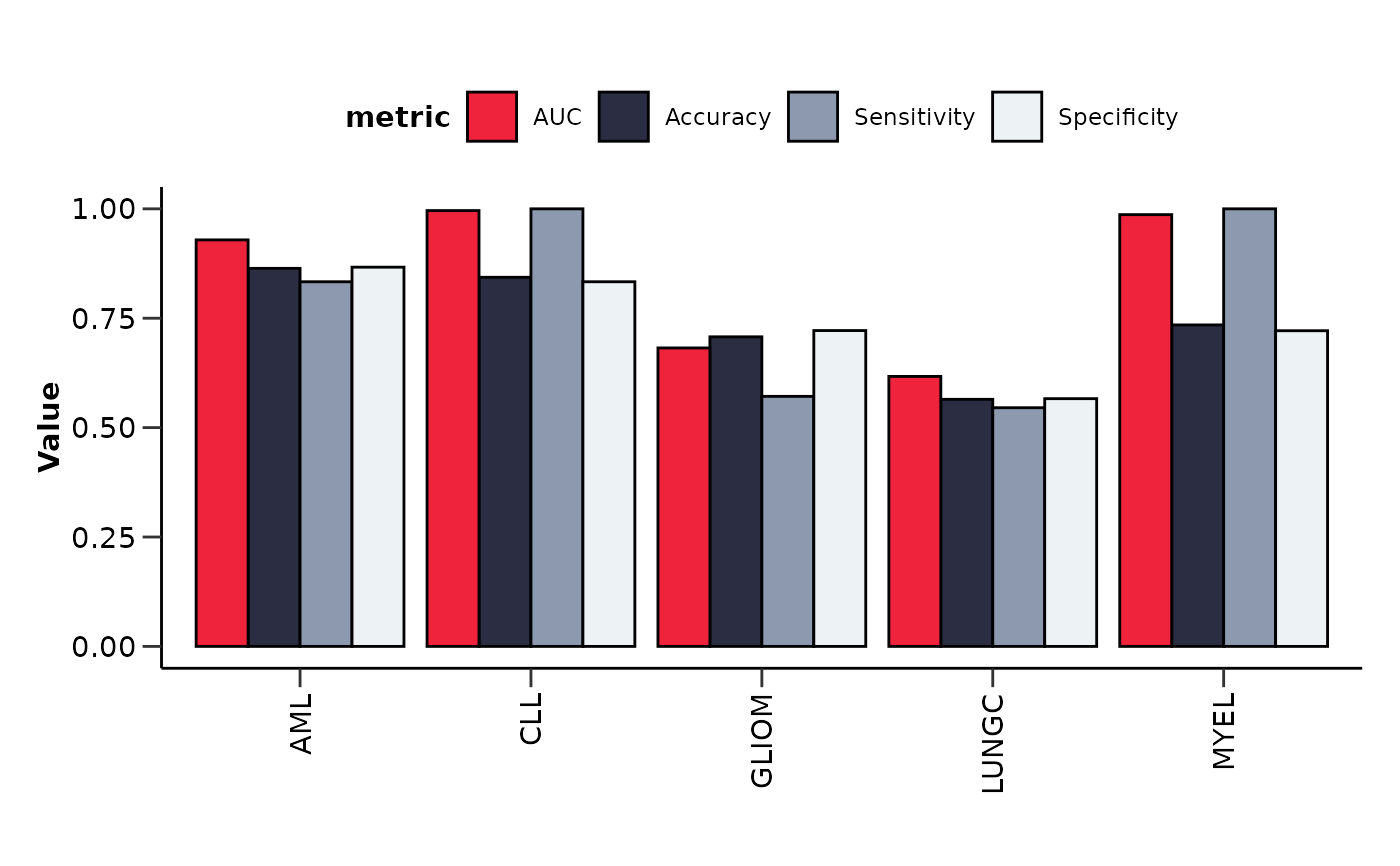 #>
#> $upset_plot_features
#>
#> $upset_plot_features
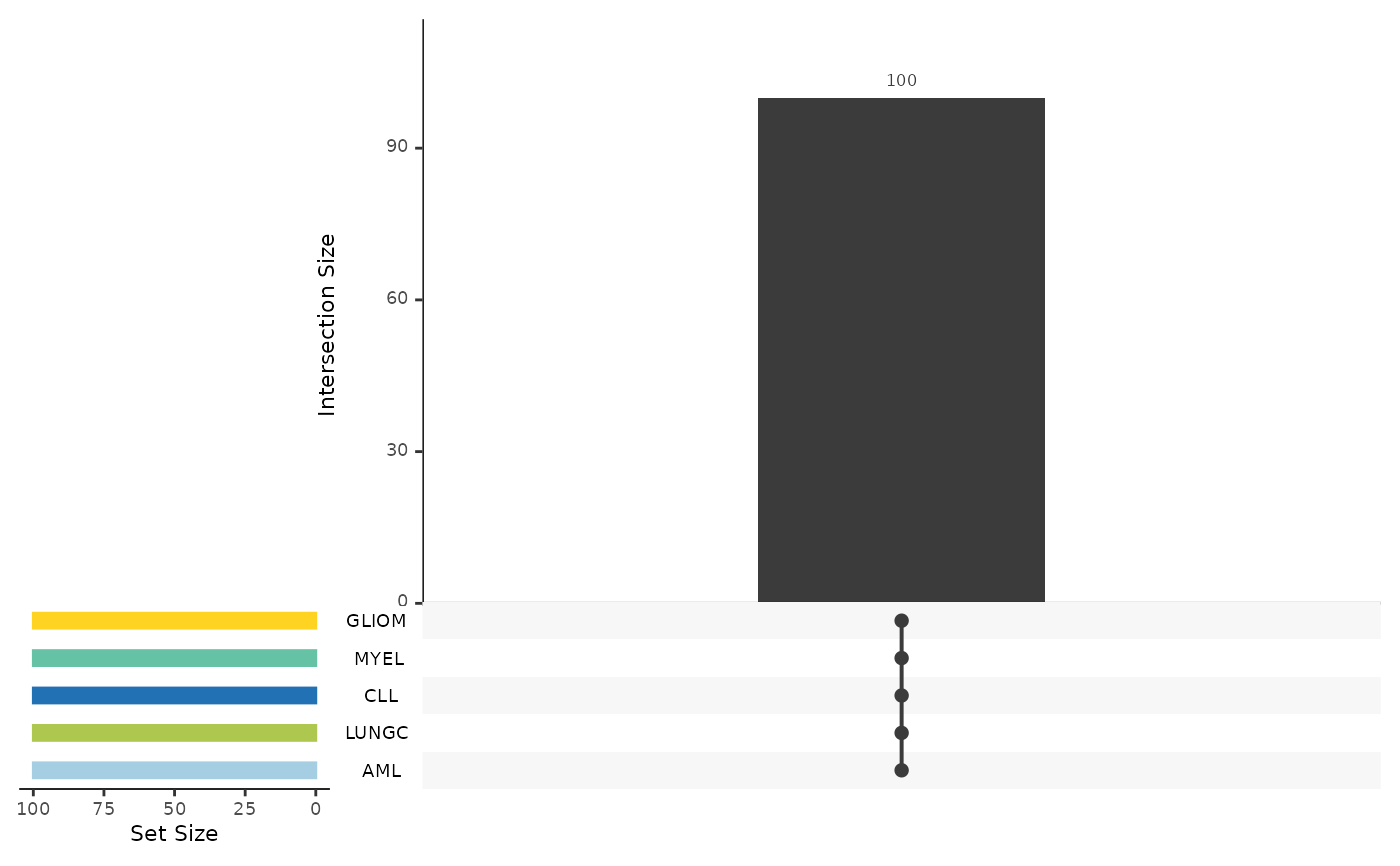 #>
#> $features_df
#> # A tibble: 100 × 3
#> Shared_in `up/down` Feature
#> <chr> <chr> <fct>
#> 1 AML&LUNGC&CLL&MYEL&GLIOM up ACAA1
#> 2 AML&LUNGC&CLL&MYEL&GLIOM up ACE2
#> 3 AML&LUNGC&CLL&MYEL&GLIOM up ACOX1
#> 4 AML&LUNGC&CLL&MYEL&GLIOM up ACP5
#> 5 AML&LUNGC&CLL&MYEL&GLIOM up ACTN4
#> 6 AML&LUNGC&CLL&MYEL&GLIOM up ACY1
#> 7 AML&LUNGC&CLL&MYEL&GLIOM up ADA2
#> 8 AML&LUNGC&CLL&MYEL&GLIOM up ADAM23
#> 9 AML&LUNGC&CLL&MYEL&GLIOM up ADAMTS13
#> 10 AML&LUNGC&CLL&MYEL&GLIOM up ADAMTS15
#> # ℹ 90 more rows
#>
#> $features_list
#> $features_list$`AML&LUNGC&CLL&MYEL&GLIOM`
#> [1] ANGPT1 ADGRG1 AMIGO2 ADAMTS16 AHCY ADA AMY2A
#> [8] APEX1 AK1 ABL1 ADAM8 ANGPTL2 APBB1IP ADH4
#> [15] ANKRD54 AMFR ADGRG2 APOM ANXA11 ALCAM ANGPT2
#> [22] ALDH1A1 ADGRE2 AARSD1 AXL ANPEP ATP6V1D AZU1
#> [29] ACTA2 AMBP APOH AGR3 ACP6 ATG4A ANG
#> [36] APP ARTN ATXN10 ACAN ARNT ATP6V1F ARHGAP1
#> [43] AOC1 AMBN AGRP ADAM15 AGRN AKR1B1 AMN
#> [50] APLP1 ACAA1 ACE2 ACOX1 ACP5 ACTN4 ACY1
#> [57] ADA2 ADAM23 ADAMTS13 ADAMTS15 ADAMTS8 ADCYAP1R1 ADGRE5
#> [64] ADM AGER AGR2 AGXT AHSP AIF1 AIFM1
#> [71] AKR1C4 AKT1S1 AKT3 ALDH3A1 ALPP AMY2B ANGPTL1
#> [78] ANGPTL3 ANGPTL4 ANGPTL7 ANXA10 ANXA3 ANXA4 ANXA5
#> [85] AOC3 AREG ARG1 ARHGAP25 ARHGEF12 ARID4B ARSA
#> [92] ARSB ART3 ATF2 ATOX1 ATP5IF1 ATP5PO ATP6AP2
#> [99] AXIN1 B4GALT1
#> 100 Levels: ACAA1 ACE2 ACOX1 ACP5 ACTN4 ACY1 ADA2 ADAM23 ADAMTS13 ... ANGPT1
#>
#>
#>
#> $features_df
#> # A tibble: 100 × 3
#> Shared_in `up/down` Feature
#> <chr> <chr> <fct>
#> 1 AML&LUNGC&CLL&MYEL&GLIOM up ACAA1
#> 2 AML&LUNGC&CLL&MYEL&GLIOM up ACE2
#> 3 AML&LUNGC&CLL&MYEL&GLIOM up ACOX1
#> 4 AML&LUNGC&CLL&MYEL&GLIOM up ACP5
#> 5 AML&LUNGC&CLL&MYEL&GLIOM up ACTN4
#> 6 AML&LUNGC&CLL&MYEL&GLIOM up ACY1
#> 7 AML&LUNGC&CLL&MYEL&GLIOM up ADA2
#> 8 AML&LUNGC&CLL&MYEL&GLIOM up ADAM23
#> 9 AML&LUNGC&CLL&MYEL&GLIOM up ADAMTS13
#> 10 AML&LUNGC&CLL&MYEL&GLIOM up ADAMTS15
#> # ℹ 90 more rows
#>
#> $features_list
#> $features_list$`AML&LUNGC&CLL&MYEL&GLIOM`
#> [1] ANGPT1 ADGRG1 AMIGO2 ADAMTS16 AHCY ADA AMY2A
#> [8] APEX1 AK1 ABL1 ADAM8 ANGPTL2 APBB1IP ADH4
#> [15] ANKRD54 AMFR ADGRG2 APOM ANXA11 ALCAM ANGPT2
#> [22] ALDH1A1 ADGRE2 AARSD1 AXL ANPEP ATP6V1D AZU1
#> [29] ACTA2 AMBP APOH AGR3 ACP6 ATG4A ANG
#> [36] APP ARTN ATXN10 ACAN ARNT ATP6V1F ARHGAP1
#> [43] AOC1 AMBN AGRP ADAM15 AGRN AKR1B1 AMN
#> [50] APLP1 ACAA1 ACE2 ACOX1 ACP5 ACTN4 ACY1
#> [57] ADA2 ADAM23 ADAMTS13 ADAMTS15 ADAMTS8 ADCYAP1R1 ADGRE5
#> [64] ADM AGER AGR2 AGXT AHSP AIF1 AIFM1
#> [71] AKR1C4 AKT1S1 AKT3 ALDH3A1 ALPP AMY2B ANGPTL1
#> [78] ANGPTL3 ANGPTL4 ANGPTL7 ANXA10 ANXA3 ANXA4 ANXA5
#> [85] AOC3 AREG ARG1 ARHGAP25 ARHGEF12 ARID4B ARSA
#> [92] ARSB ART3 ATF2 ATOX1 ATP5IF1 ATP5PO ATP6AP2
#> [99] AXIN1 B4GALT1
#> 100 Levels: ACAA1 ACE2 ACOX1 ACP5 ACTN4 ACY1 ADA2 ADAM23 ADAMTS13 ... ANGPT1
#>
#>
